Binding Properties
| 𝜈 | Molecule 1 : 1 Host | ||
| Ka = | 1442.0 | M-1 | |
| Kd = | |||
| logKa = | |||
| T | 25.0 °C | ||
| Energy | kJ mol-1 | kcal mol-1 | |||
|---|---|---|---|---|---|
| ΔG | = | -18.03 | -4.31 |
These are the specifications of the determination of the experimental results.
| Detection Method: | Direct | ||
| Assay Type: | Direct Binding Assay | ||
| Technique: | Fluorescence | ||
Detailed information about the solvation.
| Solvent System | Complex Mixture | |
| Solvents | water | 96.0 % |
| methanol | 4.0 % | |
| Additives | Citric acid mon... | |
| Trisodium citra... | ||
| pH | 6.0 |
Please find here information about the dataset this interaction is part of.
| Citation: |
D. Guo, Y. Liu, C. Li, Z. Pan, Z. Li, SupraBank 2026, A Comparative Study of Complexation of β-Cyclodextrin, Calix[4]arenesulfonate and Cucurbit[7]uril with Dye Guests: Fluorescence Behavior and Binding Ability (dataset). https://doi.org/10.34804/supra.20210928374 |
| Link: | https://doi.org/10.34804/supra.20210928374 |
| Export: | BibTex | RIS | EndNote |
Please find here information about the scholarly article describing the results derived from that data.
| Citation: |
Y. Liu, C.-J. Li, D.-S. Guo, Z.-H. Pan, Z. Li, Supramolecular Chemistry 2007, 19, 517–523. |
| Link: | https://doi.org/10.1080/10610270601145444 |
| Export: | BibTex | RIS | EndNote |
Binding Isotherm Simulations
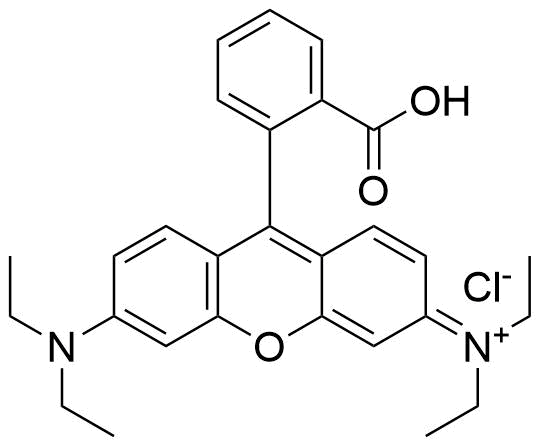
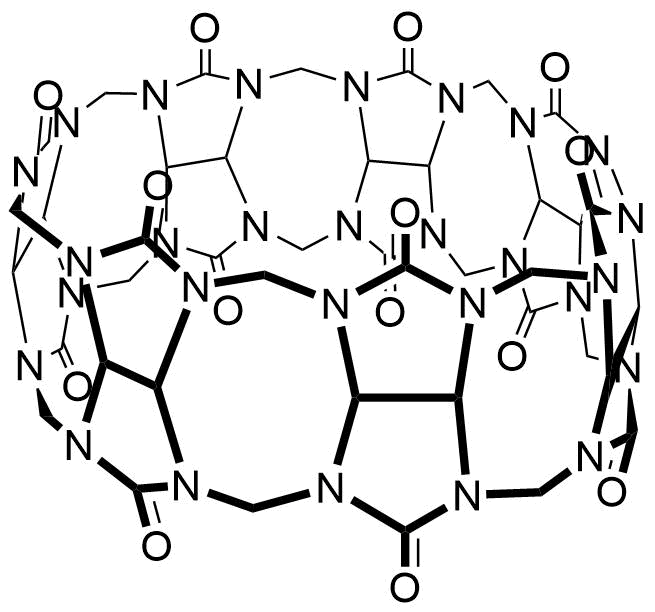
The plot depicts the binding isotherm simulation of a 1:1 interaction of Rhodamine B (0.013869625520110958 M) and CB7 (0 — 0.027739251040221916 M).
Please sign in: customize the simulation by signing in to the SupraBank.